Our aim was to perform an initial assessment of the polymorphic patterns of the PIN1 gene in patients with coronary heart disease (CHD). The PIN1-encoded protein (Pin1) suppresses eNOS-NO signaling and may impair cardiovascular function. Blood collection, DNA extraction, PCR amplification and gene sequencing were performed for thirty CHD participants living in central China, focusing on nine single nucleotide polymorphisms (SNPs). Their genetic linkages were revealed and their allele frequencies were compared with SNP data from the NCBI. Three major linkage patterns were identified: [1.rs2287839-5.rs2233682], [3.rs2233679-4.rs1077220–8.rs2287838] and [6.rs889162-7.rs2010457], suggesting correlated involvement in CHD and possible simultaneous genetic origin in ancient times. The frequencies of six SNPs are consistent with the NCBI data, while the frequencies of three SNPs (2.rs2233678, 4.rs1077220 and 9.rs4804461) are not consistent with the NCBI. Especially, the 3.rs2233679–4.rs1077220 linkage is different from other populations worldwide and may be an interesting genetic characteristic of Chinese CHD patients. Predictably, 1.rs2287839, 2.rs2233678, 3.rs2233679 and 5.rs2233682 may be strongly associated with CHD risk, although this requires future verification. The PIN1 SNP linkages lay a new genetic foundation for discovering novel molecular mechanisms of CHD and for exploring PIN1-based targeted treatment of CHD with nitric oxide regulatory therapies in clinical practice.
O objetivo deste estudo é avaliar os polimorfismos do gene PIN1 em doentes com doença arterial coronária (DAC). A proteína codificada pelo gene PIN1 (Pin1) suprime a sinalização eNOS-NO e pode afetar a função cardiovascular. Foram incluídos 30 indivíduos com DAC, residentes na China central, a quem foi recolhido sangue, extraído e amplificado o DNA por PCR. Foram estudados nove polimorfismos de nucleótidos únicos (SNPs) do gene PIN1. As suas associações genéticas foram estudadas e as frequências de alelo foram comparadas com os dados de SNPs do NCBI. Foram identificados três grandes padrões de linkage: [1.rs2287839-5.rs2233682], [3.rs2233679-4.rs107220-8.rs2287838] e [6.rs889162-7.rs2010457], sugerindo o seu envolvimento na DAC e possível origem genética simultânea histórica. As frequências de seis dos SNPs foram consistentes com os dados do NCBI, enquanto as frequências de três dos SNPs (2.rs2233678, 4.rs1077220 e 9.rs4804461) não foram consistentes com o NCBI. Especificamente, a ligação 3.rs2233679-4.rs1077220 é diferente de outras populações mundiais e poderá constituir uma característica genética específica dos doentes chineses com DAC. Previsivelmente, 1.rs2287839, 2.rs2233678, 3.rs2233679 e 5.rs2233682 podem estar particularmente associados ao risco de DAC, requerendo validação ulterior. As ligações PIN1-SNP estabelecem uma nova base genética para a descoberta de novos mecanismos moleculares da DAC e para explorar o tratamento de precisão da DAC com medicamentos reguladores do óxido nítrico, baseado nos polimorfismos do gene PIN1.
Morbidity from coronary heart disease (CHD) is increasing worldwide, posing a major threat to health. CHD correlates with obesity, a high-fat diet, smoking, drinking, and other risk factors. However, many people without these unhealthy lifestyles and dietary habits still suffer from CHD, so researchers are paying more attention to genetic differences in CHD patients.1–5
Pin1, encoded by the PIN1 gene, regulates phosphorylation of proteins and is implicated in human diseases.6–11 Our recent research predicted that (i) Pin1 may promote cardiovascular disease by suppressing endothelial nitric oxide synthase (eNOS) and production of nitric oxide (NO) in blood vessels,7 (ii) a potential glucose-Pin1-eNOS-NO pathway may lead to coronary atherosclerosis in diabetic patients,12 and (iii) serum Pin1 may be associated with CHD and hypertension.11 NO is a key vasoactive molecule that promotes vasodilation, reduces vascular sediments, and prevents atherosclerosis, but Pin1 inhibits eNOS and blocks NO bioactivity in animal and cellular experiments, and probably increases CHD risk in humans.8,9,12,13 However, no clinical research on the relationship between the PIN1 gene and CHD has been published.
PIN1 has multiple polymorphic sites that may be related to many diseases, but previous studies analyzed these polymorphic sites in isolation without paying attention to their possible genetic linkages, which inevitably complicates research work and increases the cost of analysis.10,14–19 Hence, we aim to perform an initial exploration of the genetic linkages of the nine common single nucleotide polymorphisms (SNPs) of the PIN1 gene and thus to lay a foundation for simplifying relevant research in the future. To improve analytic efficiency, we initially analyzed the PIN1 SNP linkages in CHD patients living in the city of Handan, China, and identified three typical genetically-linked SNP groups, which will facilitate future research and foster studies on the association of PIN1 and CHD (as well as potentially other relevant genetic diseases).
MethodsPatient enrollmentAll participants belong to the Han nationality and live in the city of Handan (Hebei province, China). All participants provided written information including name, gender, age, height, weight, personal information, living and dietary habits, and other details. Thirty CHD patients were randomly enrolled in the cardiology department of the Affiliated Hospital of Hebei University of Engineering. The selection criterion for CHD was at least one coronary artery with >50% stenosis. The research was approved by the Ethics Committee of the Medical College of Hebei University of Engineering, and every participant signed an informed consent form.
Blood sample preparationFor each participant, 5 ml peripheral blood was collected in the morning using anticoagulant vacuum tubes. Blood plasma and blood cells were separated by centrifugation at 3000 rpm for 10 min and were then stored at -80°C.
Polymerase chain reaction amplification of PIN1 gene fragmentsAll of the reagents used for DNA analysis were provided by Sangon Biotech, Shanghai. DNA was extracted using the Ezup column blood genomic DNA extraction kit (product number: B518253), and polymerase chain reaction (PCR) amplification was performed using Taq DNA polymerase (product number: B600090), 10× PCR Buffer (product number: B600017), and dNTP (product number: B500056). The PCR primers for analyzing the nine SNPs are shown in Table 1.
Polymerase chain reaction primers for amplifying PIN1 single nucleotide polymorphisms.
| No. | SNP sites | Primer sequences |
|---|---|---|
| 1 | rs2287839 | F: 5′-GTCAGAGTCAGGATTCCTTAACAC-3′R: 5′-ATCTACTGTCATTAGCCGGAGG-3′ |
| 2 | rs2233678 | F: 5′-AGGTCGCATAGCAAGTGTCAGT-3′R: 5′-GGAGATGAGCACCTAAGTACCC-3′ |
| 3 | rs2233679 | Same as for rs2233678 |
| 4 | rs1077220 | F: 5′-AGTGAGGGGTATGAAGGAGTGAA-3′R: 5′-AAACTGAAGGAACTCGTCCAAGA-3′ |
| 5 | rs2233682 | F: 5′-CTCTGTTCCATCACTCTGGGTTAT-3′R: 5′-AAGGCTCACCTGGGAGAAGAC-3′ |
| 6 | rs889162 | F: 5′-ATCCTTCTTGCTTCCTACTGGC-3′R: 5′-AGAATAATAAGACCCACTTCACAGG-3′ |
| 7 | rs2010457 | F: 5′-GATGATGCCAGGAAGAAAGTGAT-3′R: 5′-GTGCTCATGCTCGTGAAAGG-3′ |
| 8 | rs2287838 | F: 5′-CAGGCTACATCCAGAAGATCAAGT-3′R: 5′-AGTGCGGAGGATGATGTGGAT-3′ |
| 9 | rs4804461 | Same as for rs2287838 |
PCR products were recycled using the SanPrep Column PCR Product Purification Kit (B518141), and the DNA sequences were determined using an ABI 3730xl DNA Analyzer (Applied Biosystems).
SNP frequencies of the PIN1 gene were then compared with the US National Center for Biotechnology Information (NCBI) database (https://www.ncbi.nlm.nih.gov/snp/) as a public control.
ResultsCharacteristics of study participants with coronary heart diseaseThe clinical characteristics of the study participants are shown in Table 2. Their age distribution ranged from about 50 to 70 years, and some individuals were slightly overweight but rarely reached the level of obesity. Mean values of blood components such as triglycerides, cholesterol, bile acids, and apolipoproteins were approximately within normal ranges, indicating that these traditional risk factors had less impact on CHD development in these participants, which strengthens our belief that genetic differences may substantially influence the incidence of CHD.
Characteristics of the study population with coronary heart disease (n=30).
| Variables | Values | Remarks |
|---|---|---|
| Gender | ||
| Femalea | 30 (100%) | |
| Male | 0 (0%) | |
| Age, years | 69.17±8.58 | |
| <60 | 2 (6.7%) | |
| 50-60 | 16 (53.3%) | Major population |
| 60-70 | 6 (20.0%) | |
| >70 | 5 (16.7%) | |
| BMI, kg/m2 | 25.57±3.36 | |
| <18.4 | 1 (3.3%) | Underweight |
| 18.5-24.9 | 10 (33.3%) | Normal weight (major) |
| 25.0-29.9 | 14 (46.7%) | Overweight (major) |
| >30.0 | 1 (3.3%) | Obese |
| Blood components | Normal ranges | |
| Triglycerides, mmol/l | 1.65±0.70 | 0.48-1.88 |
| Total cholesterol, mmol/l | 5.04±1.23 | 3.35-6.45 |
| HDL cholesterol, mmol/l | 1.22±0.27 | 1.00-1.94 |
| LDL cholesterol, mmol/l | 2.77±0.93 | <3.12 |
| Total bile acid, μmol/l | 7.69±7.44 | 5.1-19 |
| Apolipoprotein A1, g/l | 1.27±0.28 | 1.0-1.6 |
| Apolipoprotein B, g/l | 1.01±0.36 | 0.6-1.1 |
| Lipoprotein(a), mg/l | 238±228 | 0-300 |
Data are presented as n (%) or mean ± standard deviation.
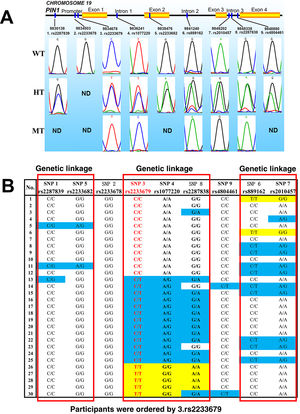
As shown in Table 1, rs2233678 and rs2233679 are close to each other on chromosome 19 and were analyzed using the same pair of primers, and similarly rs2287838 and rs4804461 were analyzed together using another pair of primers. Examples of gene sequencing are shown in Figure 1A. For six of these SNP loci, three types of genotypes were identified: wild type (WT), heterozygous type (HT), and mutant type (MT). However, only WT and HT were determined for 1.rs2287839, 5.rs2233682 and 9.rs4804461, and only the WT allele (G) was determined for 2.rs2233678.
Linkage patterns of PIN1 single nucleotide polymorphisms in the study populationFigure 1B provides clues to the genetic linkage information of PIN1 SNPs. Three typical genetic linkages were revealed: Group I, 1.rs2287839-5.rs2233682; Group II, 3.rs2233679-4.rs1077220–8.rs2287838; Group III, 6.rs889162-7.rs2010457. No significant linkage between other SNP loci was found.
Based on these genetic linkages, the impact of PIN1 on CHD can more easily be explored. In theory, examining several key SNPs, particularly 1.rs2287839, 2.rs2233678 and 3.rs2233679 in the gene promoter, will simultaneously present pathogenic information about their genetically-linked SNP loci in other exons and introns.
Single nucleotide polymorphism analysis compared with the NCBI dataIn order to better understand the racial distributions of PIN1 SNPs, we compared the SNP frequencies with the information from the NCBI's SNP database, shown in Table 3. Briefly, the frequencies of most mutant alleles are very close to the NCBI ranges, except 2.rs2233678, 4.rs1077220 and 9.rs4804461.
PIN1 allele counts (n=60) and frequencies in the study population compared to the allele information from the NCBI database.
| No. | Gene location | Potential impact | SNPs | WT alleles | MT alleles | Frequencies in NCBI |
|---|---|---|---|---|---|---|
| 1 | Promoter | Gene transcription | rs2287839 | C (57, 95.0%) | G (3, 5.0%) | G (5.6-17.0%) |
| 2 | Promoter | Gene transcription | rs2233678 | G (60, 100.0%) | C (0, 0.0%)a | C (7.7-15.1%) |
| 3 | Promoter | Gene transcription | rs2233679 | C (36, 60.0%) | T (24, 40.0%) | T (39.9-41.4%) |
| 4 | Intron 1 | RNA splicing | rs1077220 | A (36, 60.0%)a | G (24, 40.0%) | A (21.8-35.7%) |
| 5 | Exon 2 | Protein translation | rs2233682 | G (58, 96.7%) | A (2, 3.3%) | A (0.4-8.4%) |
| 6 | Intron 2 | RNA splicing | rs889162 | C (47, 78.3%) | T (13, 21.7%) | T (18.0-24.1%) |
| 7 | Intron 3 | RNA splicing | rs2010457 | A (46, 76.7%) | G (14, 23.3%) | G (29.1-39.8%) |
| 8 | Intron 3 | RNA splicing | rs2287838 | G (39, 65.0%) | A (21, 35.0%) | A (31.1-41.8%) |
| 9 | Intron 3 | RNA splicing | rs4804461 | C (58, 96.7%) | T (2, 3.3%)a | T (14.5-25.9%) |
Firstly, for 2.rs2233678, the NCBI frequency of the mutant allele C is about 7.7-15.1%, but unexpectedly we did not observe this (C: 0%, Figure 1 and Table 3) in the study participants, which implies that the homozygous WT genotype (GG, enhanced PIN1 transcription10) may elicit a higher CHD risk. In other words, people carrying the mutant allele C may face such a lower CHD risk that they were not included in our CHD participants this time, so further clinical case-control investigation will be particularly necessary for this SNP.
Secondly but also very interestingly, 3.rs2233679 and 4.rs1077220 are completely linked in the study participants, and the frequencies of their mutant alleles (T of 3.rs2233679, and G of 4.rs1077220) are both 40% (Table 3). However, although the frequency of 3.rs2233679-T (40%) is nearly the same as the NCBI data (39.9-41.4%), the frequency of 4.rs1077220-A (60%) is significantly different from the NCBI (21.8-35.7%). This inconsistency argues against complete linkage between 3.rs2233679 and 4.rs1077220 (Figure 1B). Hypothetically, this seems to indicate that the 3.rs2233679-4.rs1077220 linkage may be a specific chromosomal characteristic that distinguishes Chinese CHD patients from other racial groups, which also requires further clinical verification.
DiscussionThe first issue needing explanation is why only female patients were analyzed here. Several recent studies suggested that smoking and alcohol may interact with the PIN1 gene and have synergistic pathogenic effects in human lung cancer and in mouse cardiac cells, respectively.10,13 Therefore, only women who did not smoke or drink were enrolled, to avoid the potential influence of tobacco and alcohol on PIN1 expression. Secondly, although Figure 1B clearly indicates the potential genetic linkages, the authors do not wish to exaggerate these associations. After all, the sample size is still limited in this study, so we suggest that these preliminary conclusions should be confirmed in large-scale studies by us or by researchers in other regions of China and other countries in the future.
In theory, higher Pin1 activity in vivo could be largely attributed to highly-expressed PIN1 genotypes. Hence, the major impact of PIN1 SNPs on higher Pin1 expression as well as higher CHD risk can be predicted in the following respects:
1.rs2287839-5.rs2233682: Compared to the WT G allele of 1.rs2287839 (-5185 in the promoter), the mutant C allele may enhance PIN1 transcription by rejecting a suppressor of the PIN1 promoter.18 5.rs2233682 results in a same-sense mutation from CAG to CAA (encoding glutamine) in exon 2, but its influence on PIN1 transcription and protein translation needs to be clarified in the future.
2.rs2233678: Compared to the variant C allele (-842 in the promoter), the WT G allele enhances PIN1 transcription and is associated with several cancers.10,16,17,19,20 In this work, all of the study participants were of the GG genotype (100%, significantly higher than NCBI data), possibly indicating that the G allele may elicit a higher CHD risk.
3.rs2233679-4.rs1077220–8.rs2287838: 3.rs2233679 is another key polymorphic locus in the PIN1 promoter (-667), although its influence on Pin1 function has been somewhat controversial.16,17,21 For example, two studies recently showed that the -842C-667C haplotype has a protective effect against lung and esophageal cancer (higher Pin1 results in cancers),10,22 which hints that -667C may decrease Pin1 levels and lead to lower CHD risk. In addition, 4.rs1077220 and 8.rs2287838 may play less important roles compared to 3.rs2233679 because the two SNPs are located in the introns of the PIN1 gene.
Taken together, the presented results provide a new genetic basis for early prevention and targeted treatment of CHD. In particular, 2.rs2233678 is considered an important SNP that affects PIN1 transcription and Pin1 protein levels,10,17 whereas only the WT G allele was identified in our study population, implying that the GG genotype might be a risk factor for CHD, which necessitates further investigations in large-scale case-control studies in the future.
Typical CHD therapies such as nitrate esters, calcium channel blockers and estrogens show beneficial therapeutic efficacy by improving NO-stimulated vascular diastolic function, but recent evidence supports the hypothesis that higher Pin1 may reduce NO production and result in coronary atherosclerosis.7,8,12,13 Therefore, a possible association of the polymorphic patterns of the PIN1 gene and CHD should be considered. For example, PIN1 polymorphism may be a new genetic basis for screening high-risk CHD populations and for defining new CHD subtypes, and PIN1-based targeted prevention and treatment for CHD may be achieved via novel Pin1 inhibitors (such as juglone and small molecules that are currently being developed by structural biologists). According to the principles of pharmacogenomics,23–27PIN1 SNPs may be valuable biomarkers for optimizing the therapeutic dosages of NO-regulating drugs such as nitrates, calcium channel blockers and estrogen, in order to achieve greater efficacy as well as to minimize potential clinical side effects.
ConclusionThis work succinctly reveals the SNP profiles of PIN1 in CHD patients. To our knowledge, this is the first research that focuses on the relationship between the PIN1 gene and human CHD and is also the first explicit linkage determination of nine PIN1 SNPs in humans. This linkage analysis predicts synergistic CHD risks and may exclude redundant SNPs for more convenient, efficient and economical CHD risk assessment.
The above findings also highlight a novel genetic basis for promoting molecular mechanism studies, early prevention and targeted treatment of CHD and other Pin1-related comorbidities. These predictions need to be verified in clinical studies on CHD patients and control individuals in the near future.
FundingThis work is supported by funding from Hebei Provincial Department of Science and Technology, China (No. 182777107D).
Conflicts of interestThe authors have no conflicts of interest to declare.
The authors sincerely thank all the doctors of the department of cardiology in the Affiliated Hospital of Hebei University of Engineering.